Text Mining Tools
brat: annotation visualization and editing
Overview
Intuitive visualization and editing of text annotations is important for communicating the "meaning" of annotations and for reducing the effort of creating new annotations.
brat rapid annotation tool (brat) is a web-based tool for annotation visualization and editing. The tool is freely available and open source (MIT license).
brat supports a rich set of fully configurable annotation primitives:
- Typed text spans (e.g. entity mention)
- Binary relations (e.g. coreference)
- n-ary associations (e.g. events)
- Attributes/meta-knowledge (e.g. Negation, Speculation, etc.)
- Free-form text "notes"
These allow the tool to be applied to a wide range of text annotation tasks, including, for example, entity mention annotation, chunking, binary relation annotation, dependency syntax, and structured n-ary event annotation.
8/11/2012: Version 1.3 released
New features in v1.3 include:
- entity normalisation / linking / grounding support
- supporting embedded visualisations for web pages and web-based applications
- discontinuous text annotations
- in-built annotation tutorials and additional example corpora
- new annotation comparison functionality
- a fast, easy-to-use standalone server (experimental)
For details, please see: http://brat.nlplab.org/new-in-v1.3.html
Annotation visualization
The brat visualization functionality is based on the stav, a visualization tool created by the Tsujii laboratory of the University of Tokyo for the BioNLP Shared Task 2011. The initial focus of the tool was on the visualization of annotations for event extraction, and the visualization is thus originally designed to work also for complex structured annotations.
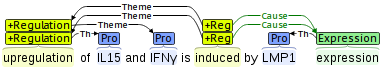
The functionality of brat has been extended to support visualization for many other annotation tasks, and brat provides also support features such as text and annotation search and concordancing in addition to visualization.
Online annotation
brat includes annotation capabilities using intuitive mouse-based editing "gestures" familiar from text editors, presentation software, and many other tools.
An annotation for a text span can be created simply by selecting that span with the mouse:
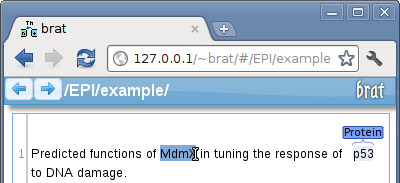
Annotations can be connected by "dragging" from one annotation to the other:
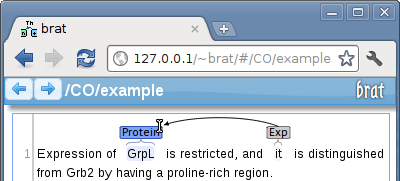
brat has been developed in close collaboration with experienced annotators working on mid-to-large-scale annotation efforts (tens of thousands of annotations), and the tool implements a full set of features for annotation support such as automatic validation of annotations against task-specific semantic constraints.
For more information, please see the brat homepage.
Contributors
The primary stav and brat developers are
- Pontus Stenetorp (Aizawa laboratory, University of Tokyo) (server development)
- Goran Topić (Tsujii laboratory, University of Tokyo) (client development)
- Sampo Pyysalo (NaCTeM and University of Manchester) (project lead)
- Tomoko Ohta (Tsujii laboratory, University of Tokyo) (quality assurance)
Acknowledgments
stav and brat development has been supported in part by
- Aizawa laboratory, University of Tokyo (PI: Akiko Aizawa)
- Tsujii laboratory, University of Tokyo (PI: Jun'ichi Tsujii)
- Grant-in-Aid for Specially Promoted Research (MEXT, Japan)
- NaCTeM and University of Manchester (PI: Sophia Ananiadou)
- UK Biotechnology and Biological Sciences Research Council (BBSRC) (reference number: BB/G013160/1)
NaCTeM is contributing to the development of brat as a collaborative open-source project.
Availability
brat is freely available with full source code under the open-source MIT license.
- brat homepage (download and installation instructions)
- brat code repository
(The older visualization tool, stav, has been superseded by brat but remains available from the stav repository.)
References
If you use brat in your work, please cite the following paper:
- Pontus Stenetorp, Sampo Pyysalo, Goran Topić, Tomoko Ohta, Sophia Ananiadou and Jun'ichi Tsujii (2012). brat: a Web-based Tool for NLP-Assisted Text Annotation. In Proceedings of the Demonstrations Session at EACL 2012. (to appear)
Other related studies:
- Pontus Stenetorp, Goran Topić, Sampo Pyysalo, Tomoko Ohta, Jin-Dong Kim and Jun'ichi Tsujii (2011). BioNLP Shared Task 2011: Supporting Resources. In Proceedings of BioNLP Shared Task 2011 Workshop. (manuscript introducing stav)
Featured News
- AI for Research: How Can AI Disrupt the Research Process?
- ELLIS Workshop on Misinformation Detection - Presentation slides now available
- 1st Workshop on Misinformation Detection in the Era of LLMs (MisD)- 23rd June 2025
- Prof. Sophia Ananiadou accepted as an ELLIS fellow
- Invited talk at the 15th Marbach Castle Drug-Drug Interaction Workshop
- BioNLP 2025 and Shared Tasks accepted for co-location at ACL 2025
- Prof. Junichi Tsujii honoured as Person of Cultural Merit in Japan
- Participation in panel at Cyber Greece 2024 Conference, Athens
- New Named Entity Corpus for Occupational Substance Exposure Assessment
Other News & Events
- CL4Health @ NAACL 2025 - Extended submission deadline - 04/02/2025
- Shared Task on Financial Misinformation Detection at FinNLP-FNP-LLMFinLegal
- FinNLP-FNP-LLMFinLegal @ COLING-2025 - Call for papers
- Keynote talk at Manchester Law and Technology Conference
- Keynote talk at ACM Summer School on Data Science, Athens








