Entity normalisation
HYPHEN Method
We have applied an automatic normalisation method (HYPHEN) to link two of our three NE types, i.e., Pharmacological_substance and Disorder, with concept IDs in terminological resources.
- Pharmcological_substance entities are linked to MeSH IDs
- Disorder entities are linked to UMLS CUIS that correspond to SNOMED-CT concepts
HYPHEN is a hybrid method that it employs a pipeline of different techniques to generate variations of the original NE mention (mostly based on systematic syntactic and semantic variations of the original mention) and tries to match generated variants against existing variants listed in the target terminological resources. HYPHEN employs the following six individual techniques:
- Acronym and abbreviation expansion and context-sensitive disambiguation (e.g. elevated ICP -> elevated intracranial pressure)
- Conversion of plural forms to singular (e.g., thrombi-> thrombus)
- Generation of English equivalents of Neoclassical compounds (e.g. hyperglycemia -> high blood sugar)
- Generation of Neoclassical equivalents of English terms (e.g., thyroid englargement -> thyromegaly)
- Generation of syntactic variants (e.g., abdomen pain -> painful abdomen)
- Generation of synonyms (e.g., cardiac asystole -> cardiac arrest)
The processes are applied in the order shown above, based on the results of experiments to determine the optimal ordering. The output of each process is passed as input to the next technique in the sequence. This can increase the accuracy of normalisation, since multiple transformations are sometimes necessary to allow mapping to the terminological resource, e.g. ,hypertensive eyes -> (singular) -> hypertensive eye -> (syntactic) -> eye hypertension -> (Neoclassical) -> ocular hypertension [UMLS: C0028840]. The pipeline is terminated as soon as one of the techniques generates a variant that can be matched in the terminological resource.
Thompson, P. and Ananiadou, S. (2018). HYPHEN: A flexible, hybrid method to map phenotype concept mentions to terminological resources. Terminology, 24(1), 91-121.
HYPHEN Evaluation
To demonstrate that HYPHEN performs well when normalising NEs to these resources, we applied it to relevant gold standard annotated corpora that include mappings to our chosen resources, i.e, the ShaRE/CLEF corpus in which disorders are normalised to UMLS concept unique identifiers (CUIs) associated with SNOMED-CT concepts, and the Biocreative V CDR corpus in which chemicals are normalised to MESH IDs. In each case, we compare the performance of HYPHEN to two different baselines.
- Dictionary lookup (i.e., exact matching of NEs against variants in the respective resource).
- Results of applying the UMLS Norm program, which transforms terms in various ways, such as removing word inflections, stop words and re-ordering words alphabetically to try to match terms occurring in text against variants listed in a resource. We restrict matching to concepts that occur in the target terminological resource.
| Method | Precision | Recall | F-Score |
|---|---|---|---|
| Dictionary lookup | 90.34% | 59.51% | 71.76% |
| UMLS Norm | 84.82% | 66.65% | 74.65% |
| HYPHEN | 82.28% | 86.90% | 84.53% |
| Method | Precision | Recall | F-Score |
|---|---|---|---|
| Dictionary lookup | 98.07% | 78.78% | 87.36%% |
| UMLS Norm | 96.11% | 82.82% | 88.97% |
| HYPHEN | 94.64% | 95.91% | 96.27% |
Statistics of applying HYPHEN to PHAEDRA
In Tables 3 and 4, we report on the number of disorder and pharmacological substance NEs in the PHAEDRA to which HYPHEN is able to assign IDs corresponding to concepts in either SNOMED-CT or MeSH. We compare this figure to the number of ID assignments achieved by applying the baseline methods, which is considerably lower.
| Method | Total terms normalised | % total terms normalised |
|---|---|---|
| Dictionary lookup | 2292 | 56.26% |
| UMLS Norm | 2711 | 64.64% |
| HYPHEN | 3094 | 75.94% |
| Method | Total terms normalised | % total terms normalised |
|---|---|---|
| Dictionary lookup | 6064 | 74.88% |
| UMLS Norm | 6562 | 81.03% |
| HYPHEN | 7156 | 88.37%% |
Visualisation of concept IDs in brat
When visualising the annotated corpus, information about disorders and pharmacological substances that have been automatically linked to concept IDs by HYPHEN can be viewed by hovering the mouse over the relevant entity. Should normalisation have been carried out for the term, then the following information will be displayed:
- Concept ID assigned to the entity. This appears right aligned underneath the horizontal line.
- The heading term or prefered terms/syonyms for the entity. These are preceded by either "MESH" ot "SNOMED_CT" in bold, depending on whether the entity is a pharmacolgical substance or a disorder.
For example, Figure 1 shows that the disorder "prostate cancer" has been assigned the UMLS CUI C0376358, while in Figure 2, "heparin" has been assigned the MeSH ID D006493.
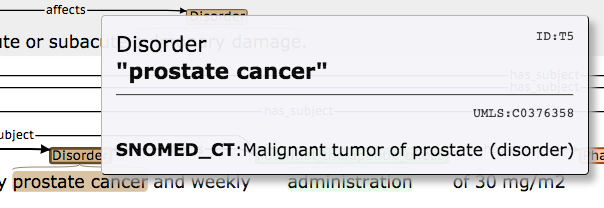 Figure 1: Display of concept ID information for disorder
Figure 1: Display of concept ID information for disorder
 Figure 2: Display of concept ID information for pharmacological substance
Figure 2: Display of concept ID information for pharmacological substance

